| Jump to section |
Introduction | Dataset and Workflow | Tour Guide | Web Sources Used | References |
Introduction
Chicken2K archives 1986 genomes representing 170 populations from 35 countries, and provides information for researchers to understand the genomic diversity across wild junglefowl and chicken populations. The database is designed with multi-functions and interactive graphical interface to visualize population genomic analyses including genetic affinities, population structure, runs of homozygosity, linkage disequilibrium decay, effective population size change, selective signals, etc. The LoucsZoom, Echarts, and d3 components are used in order to embed data in an interactive chart interface. Users can intuitively get the analysis results. Chicken2K will be provided ongoing support and long-term maintenance.
Dataset and Workflow
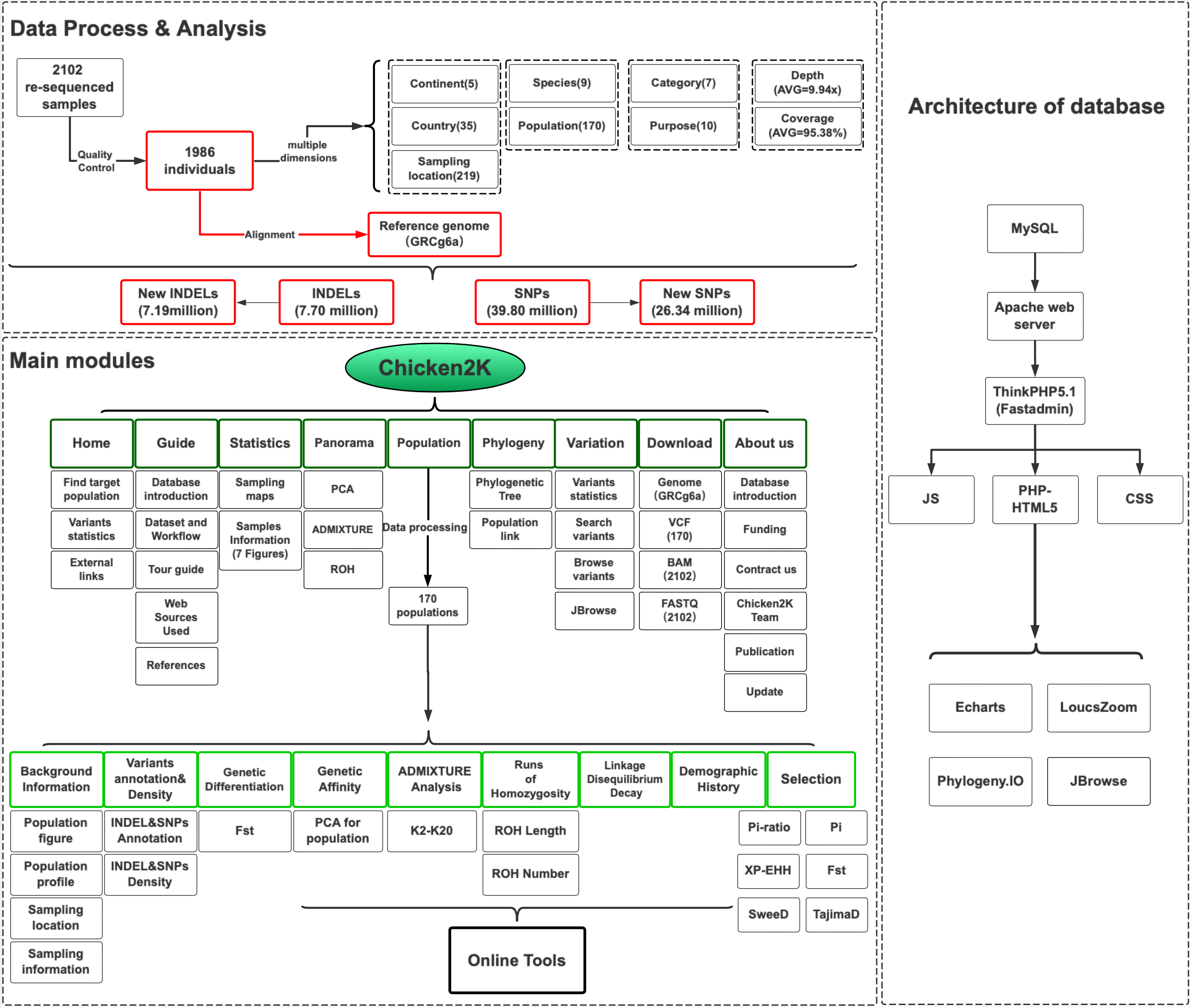
We collected a total of 2102 re-sequenced genomes for domestic chicken and wild junglefowl samples (up to July 31th, 2019). The newest chicken genome assembly GRCg6a is used as reference. After quality control and data filtering, 1986 genomes are maintained in the population genomic analyses. The sequencing depth is >4X. And the genome coverage is >85%. A total of 39.8 million high-quality SNPs and 7.7 million small INDELs have been identified. The workflow was described as following figure.
Tour Guide
We used the Serama (Wang, et al. 2017. Mol Evol Biol. DOI: 10.1093/molbev/msx227), the smallest breed of chicken, for showing the use of Chicken2K.
User can download the tour guide here
Web Sources Used
Apache web server ( http://www.apache.org);
Bootstrap ( https://getbootstrap.com);
d3 components ( https://d3js.org);
Echarts ( https://echarts.apache.org/zh/index.html);
JavaScript ( https://www.javascript.com);
JBrowse ( http://jbrowse.org);
JQuery ( https://jquery.com);
LoucsZoom ( http://locuszoom.sph.umich.edu);
MySQL( https://www.mysql.com);
phylogeny.io ( https://github.com/oist/phylogeny-io);
ThinkPHP5.1 ( http://www.thinkphp.cn)
ADMIXTURE, https://speciationgenomics.github.io/ADMIXTURE/
ANNOVAR, http://annovar.openbioinformatics.org/en/latest/
Btrim, http://graphics.med.yale.edu/trim/
BWA-MEM, http://bio-bwa.sourceforge.net/
EIGENSOFT, https://github.com/DReichLab/EIG/
GATK, https://software.broadinstitute.org/gatk/
GCTA, https://gump.qimr.edu.au/gcta
HPC, https://github.com/anindya028/HPC
Picard, https://broadinstitute.github.io/picard/
PLINK, http://zzz.bwh.harvard.edu/plink/
PopLDDecay, https://github.com/BGI-shenzhen/PopLDdecay
RapidNJ, https://birc.au.dk/Software/RapidNJ/
SAMtools, http://samtools.sourceforge.net/
SMC++, https://github.com/popgenmethods/smcpp
SweeD, https://github.com/alachins/sweed
Vcftools, http://vcftools.sourceforge.net/
References
[1] 国家畜禽遗传资源委员会. 中国畜禽遗传资源志-家禽志(Animal Genetic Resources in China – Poultry)[M]. 中国农业出版社, 2011.
[2] Joseph B, Catrin R, Janet D, Mark H, Andy C (2012). The Chicken[M]. IBSN-13: 978-1-937994-03-7.
[3] Carol E (2007). Storey’s Illustrated Guide to Poultry Breeds[M]. IBSN: 978-1-58017-667-5.
http://www.fao.org/dad-is/en/
http://www.poultryclub.org
https://heritagepoultry.org
http://eng.agraria.org
https://domesticanimalbreeds.com
https://www.domesticforest.com
https://poultrykeeper.com
https://poultrycaresunday.com